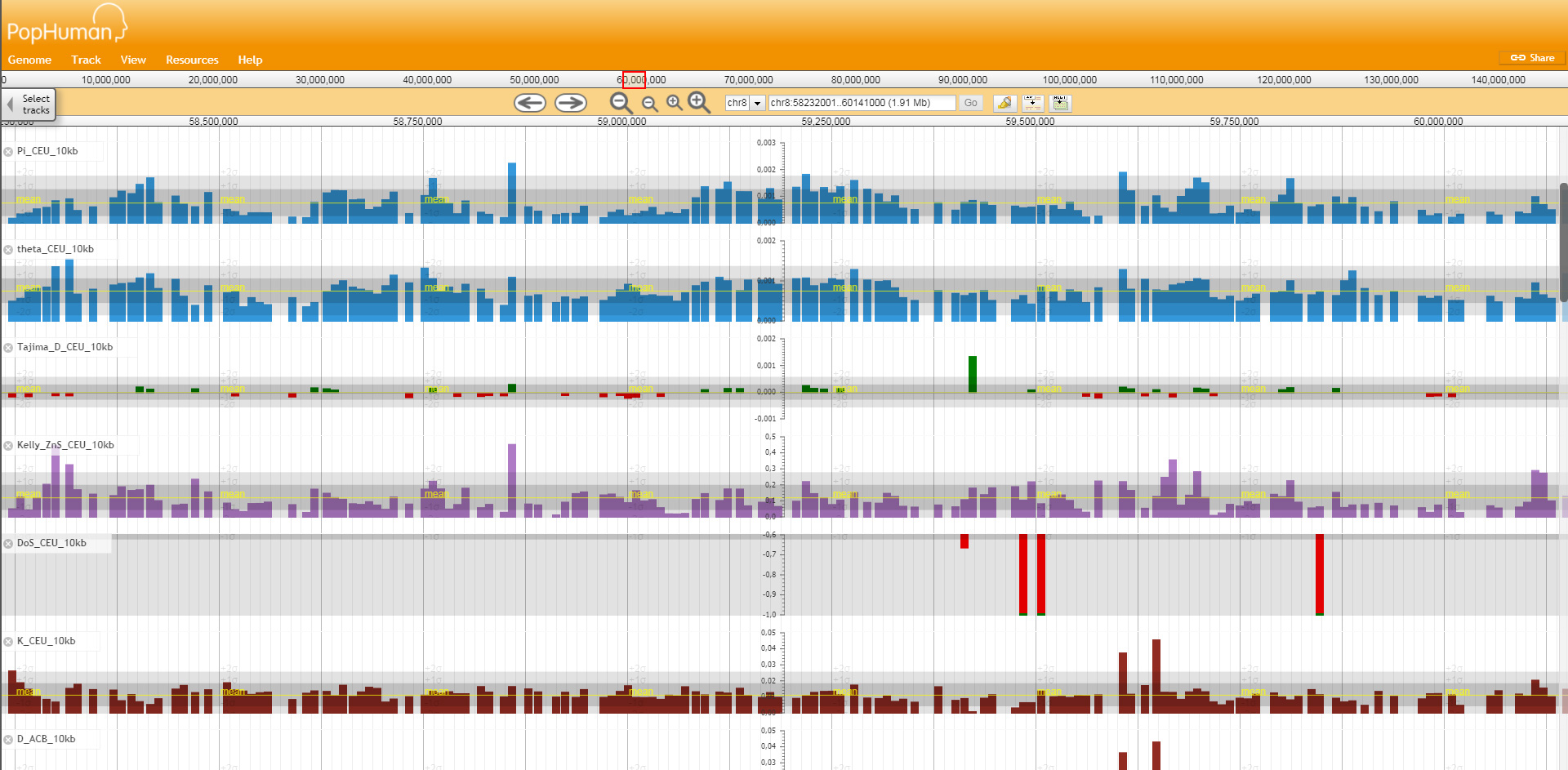
New Portal Allows Users to Navigate Thousands of Data on Human Genetic Diversity. Researchers from the Bioinformatics of Genomic Diversity group at the Universitat Autònoma de Barcelona (UAB), in collaboration with scientists from the Institute of Evolutionary Biology (IBE), a centre belonging to the CSIC and the Pompeu Fabra University (UPF), created the PopHuman browser, the largest available inventory of measures of genetic diversity along the human genome.
The PopHuman browser allows users to detect episodes of genetic selection which have left a mark on the human genome, such as those related to the establishment of agriculture in European and Asian populations some 10,000 years ago. The international project 1000 genomes provides information on 2504 complete genomic sequences from 26 populations around the world. It is the most extensive global human genetic variation data set to date, with more than 84 million genetic differences catalogued. This resource allows the exhaustive search of episodes of natural selection which have taken place in the human lineage and have left their print on the genome.
Human genetic variation is the set of genetic differences which distinguish our genomes, either among individuals within a population or between populations. There are no two genetically identical humans. On average, each of us is more than 99% genetically identical to any other person, while the less than 1% of genetic variants distinguishing our genomes are present in different frequencies in different human populations. The geographically and ancestrally more distant populations tend to differ more, although it is the differences between individuals within populations, especially in the African ones, the most important quantitatively in the set of the human genetic variation.
The study of the human genetic variation has significant evolutionary and medical applications. It has allowed to determine the origin of humanity in sub-Saharan Africa hundreds of thousands of years ago and subsequent human migrations, and it also helps in understanding how certain genetic variants have contributed to the adaptation of human populations to their environment in the whole planet. The description and explanation of the evolutionary forces controlling human genetic variation within and between populations is thus a fundamental goal of population genetics.
In an article published recently in the prestigious journal Nucleic Acids Research, the UAB and IBE researchers presented an exhaustive inventory of genetic diversity measures along the human genome from the data of the project 1000 genomes. This inventory captures the evolutionary properties of the sequences, and it is available to researchers through the PopHuman genome browser, an online portal allowing users to navigate the results through a graphical interface. PopHuman offers the most complete catalogue of estimates of human genetic variation so far in terms of number of genomes and populations analysed, and incorporates the genetic changes between the human and the chimpanzee genomes.
In this way, PopHuman allows to detect from relatively recent selective episodes, such as those related to the establishment of agriculture in European and Asian populations some 10,000 years ago, to older episodes which have recurrently occurred throughout the past million years, such as the one observed in the region of the PRM1 gene, involved in the compaction of sperm DNA. Researchers are convinced that PopHuman will become the starting point for any exhaustive analysis of the different selective episodes which have taken place in the human lineage and have left their mark on the genome.
For more information: PopHuman browser – http://pophuman.uab.cat
Reference article: Sònia Casillas, Roger Mulet, Pablo Villegas-Mirón, Sergi Hervas, Esteve Sanz, Daniel Velasco, Jaume Bertranpetit, Hafid Laayouni, Antonio Barbadilla. “PopHuman: the human population genomics browser.” 2018. Nucleic Acids Research, 46(D1):D1003-D1010. https://doi.org/10.1093/nar/gkx943. Impact Factor: 10.162.